Nanopore sequencing is a third generation approach used in the sequencing of biopolymers- specifically, polynucleotides in the form of DNA or RNA. Using nanopore sequencing, a single molecule of DNA or RNA can be sequenced without the need for PCR amplification or chemical labeling of the sample.
Oxford Nanopore Technologies Limited is a UK-based company which is developing and selling nanopore sequencing products for the direct, electronic analysis of single molecules.
A nanopore is a very small hole
A nanopore is a nano-scale hole. In its devices, Oxford Nanopore passes an ionic current through nanopores and measures the changes in current as biological molecules pass through the nanopore or near it. The information about the change in current can be used to identify that molecule.
Holes can be created by proteins puncturing membranes (biological nanopores) or in solid materials (solid-state nanopores).Learn more about biological and solid-state nanopores

How does nanopore DNA/RNA sequencing work?
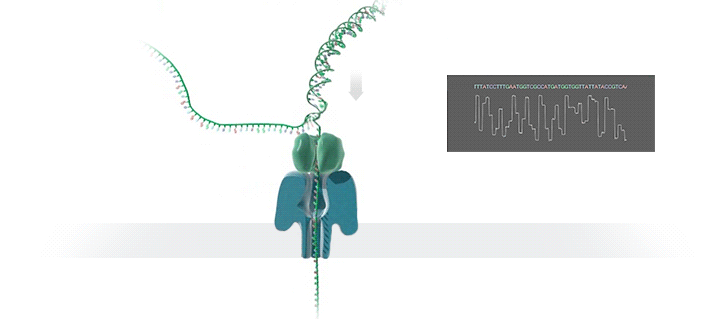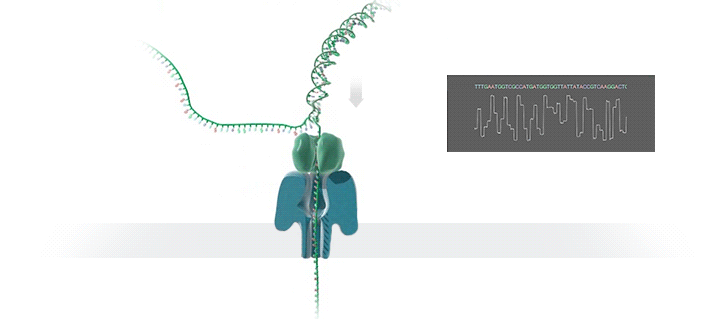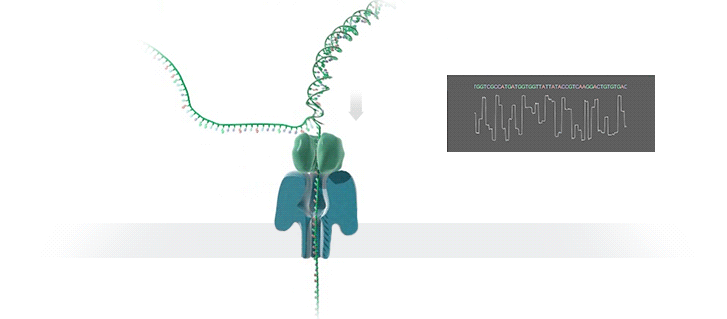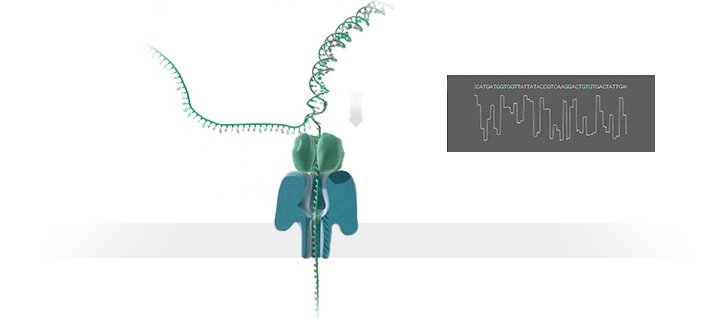
A protein nanopore is set in an electrically resistant polymer membrane. An ionic current is passed through the nanopore by setting a voltage across this membrane. If an analyte passes through the pore or near its aperture, this event creates a characteristic disruption in current (as shown in the diagram below). Measurement of that current makes it possible to identify the molecule in question.
DNA sequencing
A strand of DNA is passed through a nanopore. The current is changed as the bases G, A, T and C pass through the pore in different combinations.
Making scalable nanopore devices
Oxford Nanopore devices are fully scalable, based around a core sensing unit of a nanopore set in an arrayed sensor chip and used with a bespoke Application-Specific Integrated Circuit (ASIC) which controls and measures the experiments.

Nanopore
A protein nanopore is set in an electrically-resistant polymer membrane.

Array of microscaffolds
Each microscaffold supports a membrane and embedded nanopore. The array keeps the multiple nanopores stable during shipping and usage.

Sensor chip
Each microscaffold corresponds to its own electrode that is connected to a channel in the sensor array chip. Sensor arrays may be manufactured with any number of channels.

ASIC
Each nanopore channel is controlled and measured individually by the bespoke ASIC. This allows for multiple nanopore experiments to be performed in parallel. More than one ASIC may be included in a device and Oxford Nanopore is building ASICs of different sizes for different purposes.Learn how nanopore sensing can scale
MinION flow cell


PromethION flow cell


MinION
- Pocket-sized
- Up to 512 nanopore channels
- Simple 10-minute sample prep available
- Many publications illustrate broad usage
- Commercially available

GridION
- Multiple sequencing devices, one compute module
- Use up to five MinION Flow Cells at a time
- Benchtop processor capable of handling high data volumes in real time
- Rapid, real-time applications such as Read Until

PromethION
- Benchtop system
- Up to 48 flow cells of 3,000 nanopore channels each
- Simple 10-minute sample prep available
- Flow cells may be run individually or concurrently
Nanopore sequencing offers scientific researchers:
- Sequence any DNA/RNA fragment length from short to ultra-longCharacterise more genetic variation, versatile to broad applications
- Direct sequencing of native DNA/RNAGenerate content-rich data, including methylation
- Data available in real timeRapid insights, and analyses that can respond to results in real time
- Scalable from portable devices to ultra-high throughput desktop devicesSequence anything, anywhere
- No capital investment requiredAccessible and cost effective
- Simple & rapid, or automated, library prepEasy to use and versatile
https://nanoporetech.com/sites/default/files/s3/literature/product-brochure.pdf
