A supercomputer is a computer with a high level of performance as compared to a general-purpose computer. The performance of a supercomputer is commonly measured in floating-point operations per second (FLOPS) instead of million instructions per second (MIPS)
Innovative Computing Lab with the University of Tennessee
Fugaku – exascale supercomputer, at the RIKEN Center for Computational Science in Kobe, Japan.
It’s one of the top in performance in computational methods for industrial use, artificial intelligence applications and big data analytics.
Fugaku performed over 442 quadrillion computations per second, around three times faster than the Summit system developed by the U.S. Oak Ridge National Laboratory.
The Japanese supercomputer, which was jointly developed with Fujitsu Ltd. at Riken’s facility in Kobe, forms a key foundation for powerful simulations used in scientific research and the development of industrial and military technologies.

IBM-built supercomputers employing Power9 CPUs and NVIDIA Tesla V100 GPUs. Oak Ridge National Laboratory’s Summit system holds 148.6 petaflops.
AI breakthrough with the NVIDIA® Tesla® V100 Tensor Core, powered by NVIDIA Volta architecture. The V100 comes in 16 and 32GB configurations, and offers the performance of up to 100 CPUs in a single GPU.

Sierra system – at Lawrence Livermore National Laboratory system supports 94.6 petaflops.
IBM-built Sierra supercomputer provides four to six times the sustained performance and five to seven times the workload performance of its predecessor Sequoia, with a 125 petaFLOP/s peak. At approximately 11 megawatts, Sierra is also about five times more power efficient than Sequoia. Sierra combines two types of processor chips—IBM’s Power 9 processors and NVIDIA’s Volta graphics processing units (GPUs). It is designed for more efficient overall operations and is a promising architecture for extreme-scale computing.

Processor Architecture
IBM Power and NVIDIA Volta processors
Operating System
RHEL
Cluster/System Usage
Capability parallel computing
Processor Clock Rate/Process Clock Speed
3.1 GHz
Nodes
4,320
Cores per Node
44
Total Cores
190,080
Memory per Node
256 GB (CPU), 64 GB (GPU)
Total Memory
1.38 PB
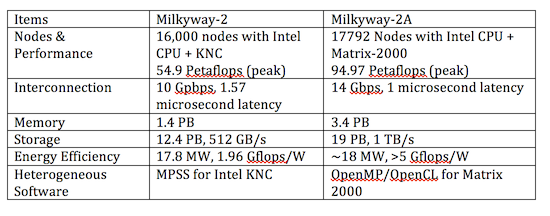
Tianhe-2A – This supercomputer, also known as Milky Way-2A, is located at the National Supercomputer Center in Guangzhou.
Sunway TaihuLight supercomputer, with an HPL mark of 94.97 petaflops. TaihuLight was developed by China’s National Research Center of Parallel Computer Engineering & Technology (NRCPC) and is installed at the National Supercomputing Center in Wuxi. It is powered exclusively by Sunway’s SW26010 processors.
NASA Supercomputers Power COVID-19 Research
NASA is flexing its supercomputing muscle to help crack some of the most pressing questions surrounding COVID-19, from basic science on how the virus interacts with cells in the human body to genetic risk factors to screening for potential therapeutic drugs.
In addition to its support of Earth, planetary, aerospace, heliophysics and astrophysics projects, the agency’s supercomputer at NASA’s Ames Research Center in California’s Silicon Valley, also has an allocation of time on it reserved for national priorities.
NASA has joined a consortium of institutions that is pairing up supercomputing resources with proposals for using high-end computing power for COVID-19 studies. The effort was organized by the White House Office of Science and Technology Policy and includes industry partners IBM, Hewlett Packard Enterprise, Amazon, Microsoft and others, as well as the Department of Energy’s National Labs, the National Science Foundation, and many universities. The consortium is supporting 64 projects and is open to new proposals. To date, four projects have been matched to NASA.
“This is not NASA’s normal work, but we have the supercomputers and the expertise to help researchers working on COVID-19 get the most out the supercomputing power,” said Tsengdar Lee, program manager for NASA’s High End Computing Program at NASA Headquarters in Washington, D.C.
Supercomputers are suited for processing large amounts of data. For NASA’s usual projects this means simulating the movements of air masses and water around the planet to study Earth’s climate, hunting for exoplanets, studying the behavior of black holes, or designing aeronautic or aerospace vehicles. Each piece of these very large puzzles is guided by certain physical and chemical laws in their interactions and relationships with other components. Zooming in to the atomic level to study the coronavirus is no different. Each molecule and cell moves and reacts based on physics and chemistry, which makes simulation a powerful tool for understanding the coronavirus.
“This is work we are pleased to support,” said Piyush Mehrotra, division chief of the NASA Advanced Supercomputing Division at Ames. “Our team will do whatever it takes to support the research projects so that we can understand and fight this disease as quickly as possible.”

Pleiades’ rack-based architecture allows NASA to continually increase the system’s computing capability through hardware upgrades without needing to expand its physical footprint. The current configuration of Pleiades is nearly 15 times more powerful than it was when the system was originally installed in 2008.Credits: NASA’s Ames Research Center
Identifying Genetic Risk Factors for Acute Respiratory Distress Syndrome
Ames’ supercomputing power is being drafted to look at the genetic risk factors associated with COVID-19 patients developing Acute Respiratory Distress Syndrome, or ARDS. ARDS is a complication of COVID-19 that occurs when the disease causes fluid to build up in the lungs, which often requires a ventilator to help patients breathe.
The same NASA researchers who apply their expertise to studying how biology changes in space are studying these genetic variations. They are partnering with health care provider, Northern California Kaiser Permanente, for their COVID-19 study.
Principal investigator Viktor Stolc and co-investigator David Loftus, both with Ames’ Space Biosciences Division, are leading a team to do genetic sequencing of patients at various stages of the COVID-19 disease who both do and do not develop ARDS.
“Not all patients are equally at risk of developing ARDS,” said Loftus. By comparing the groups and analyzing the relationships between genes and COVID-19 outcomes, the team hopes to identify genetic risk factors that make people predisposed to developing ARDS, he said. With these risk factors identified, clinicians can potentially identify patients at higher risk of complications before they become severe, and improve their care.
Processing 3D Molecular Geometry to Search for Possible Drug Therapies
A team led by Rafael Gomez-Bombarelli at the Massachusetts Institute of Technology (MIT) in Cambridge, Massachusetts, is harnessing the Ames supercomputer to train an algorithm to detect potential molecules that could inhibit the novel coronavirus from attacking cells. The machine learning algorithm will be taught with 300,000 molecules that laboratory experiments have shown to be active or not against SARS, the related coronavirus that had a deadly outbreak in 2003.
Because of the biological similarities between SARS and the novel coronavirus, “Once the algorithm is trained with the SARS data, it’s relatively easy to tweak the algorithm with very small modifications to see if the molecules would work against COVID,” said Gomez-Bombarelli.
The MIT software that will run on the supercomputer produces new 3D models of the molecules from their known chemical compositions. This allows the computer to represent the molecule more accurately, so that, when presented with a new molecule it hasn’t seen before, it can better predict whether it will bind with the novel coronavirus.
The trained algorithm can then look at a catalogue of existing therapeutic drugs at the molecular level to find those that contain molecules that are likely to be biologically active against the novel coronavirus. Drugs that have already been approved by the U.S. Food and Drug Administration or similar agencies worldwide as safe for patients are the fastest route to finding drugs that can be used soonest in hospitals to treat the novel coronavirus.

This illustration, created at the Centers for Disease Control and Prevention (CDC), reveals ultrastructural morphology exhibited by coronaviruses. Note the spikes that adorn the outer surface of the virus, which impart the look of a corona surrounding the virion, when viewed electron microscopically. A novel coronavirus, named Severe Acute Respiratory Syndrome coronavirus 2 (SARS-CoV-2), was identified as the cause of an outbreak of respiratory illness first detected in Wuhan, China, in 2019. The illness caused by this virus has been named coronavirus disease 2019 (COVID-19).Credits: CDC
Understanding the Novel Coronavirus’s Protein Shell
Understanding how the novel coronavirus uses its spiked proteins on its exterior to enter cells will better help researchers to determine what drugs or therapies may be most effective against it. Once in the cell, the novel coronavirus hijacks the cell’s functions to replicate itself, so more virus can spread through the body.
“It’s a fairly complex process, because the spike protein is in what’s called a pre-fusion state, the state before it fuses to specific receptor molecules on the cell membrane,” said principal investigator Michael Peters from Virginia Commonwealth University, in Richmond. Like many viruses, it goes through a transformation process, and the team wants to understand in detail how the spike protein behaves, he said.
Peters and his colleagues are using the Ames supercomputing resources to simulate each complex molecule at the atomic level, that is, the carbon, oxygen, nitrogen and hydrogen atoms that make up the molecules of the spike protein and its receptor. Following each atom’s movements in concert allows the study of conformational changes — changes in the overall molecule’s shape — that naturally take place with these complex molecules and their interactions.
The goal is to understand the fundamental processes the novel coronavirus spike protein uses to gain entry into cells. With this information, researchers will be able to narrow down the list of drug targets even faster to find effective treatments, Peters said.
Identifying COVID-19 Related Biomarkers
Once in the body, the novel coronavirus interacts with many variables and can potentially lead to vastly different outcomes, from relatively mild cases to severe lung conditions that require intensive care, or other organ failure. A team of researchers, called the COVID-19 International Research Team (COV-IRT), co-led by principal investigator Afshin Beheshti at Ames are analyzing the RNA sequences from patient nose swabs that include genetic material from the patient, the novel coronavirus and naturally occurring bacteria that live in the human body.
Their goal is to use the analytical power of the supercomputer to identify RNA sequences from the human and bacteria sources that interact with the RNA from the coronavirus and lead to severe disease outcomes.
Beheshti is particularly interested in the role of microRNAs—22-nucleotide fragments of RNA that can each control the expression of up to 500 genes.
“Viruses might hijack a microRNA to evade the immune system and replicate,” Beheshti said. “The idea is that the virus would take in these tiny fragments and trick the body into thinking that the virus is not a foreign object.”
In an earlier analysis on a public dataset from Wuhan, China, the team already identified a potential microRNA that’s activated by the novel coronavirus that they’ve passed on to colleagues to study with laboratory experiments.
To learn more about the NASA Ames Advanced Supercomputing Division, visit: https://www.nas.nasa.gov/
By Ellen Gray
NASA’s Earth Science News TeamLast Updated: Aug. 21, 2020Editor: Ellen Gray
